Technology
Leveraging advances in computation to transform the
design, discovery, and development of therapeutic antibodies
In Silico Design
The next generation of therapeutic antibodies are being designed by computers. At EVQLV, we continuously expand our capabilities with in silico, de novo antibody discovery to transform the pace, cost, and scale of drug development.
Our technology enhancements promise to facilitate the discovery of antibodies against challenging therapeutic targets, while saving precious resources that our partners can redeploy in pursuit of the development of additional novel therapeutics.
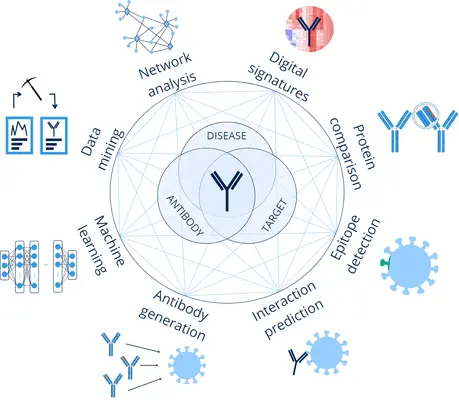
The Abtique Platform
Abtique is powered by HyperBind, a sequence-driven antibody design platform that combines validated learning models with human oversight. In validation studies, HyperBind ranked tens of millions of antibody–antigen sequence pairs in under 48 hours, accelerating candidate discovery while maintaining reproducibility.
In validation studies, HyperBind generated 20 high-affinity antibodies (KD ≤ 100 nM) against a challenging GPCR target, including 3 candidates below 10 nM. These outcomes demonstrate that sequence-based design can achieve binding strengths comparable to therapeutic-grade antibodies.
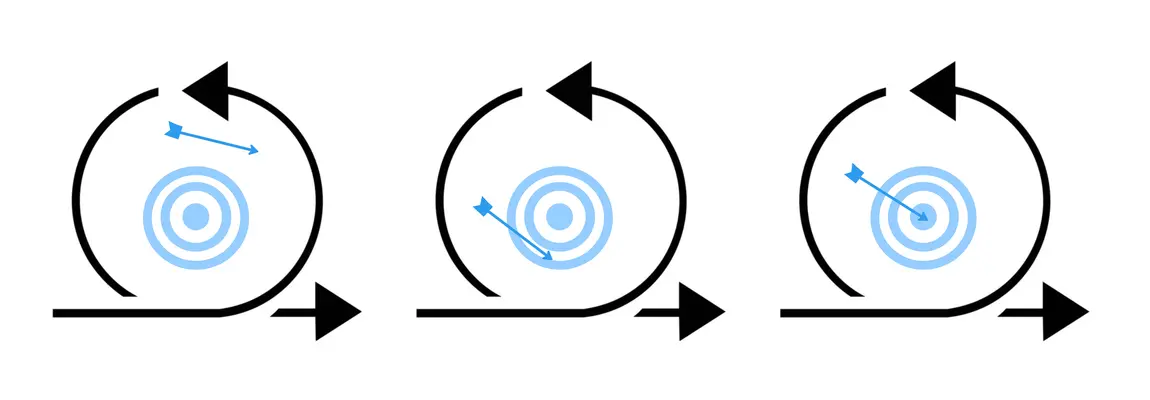
Smarter Iterations
Traditional drug discovery methods require extensive cycles of guesswork followed by time-consuming rounds of engineering and assays to identify lead-ready candidates. This process is not only long, it is expensive, risky, and failure-prone. The EVQLV platform continuously demonstrates the capacity to select viable therapeutic candidates, minimizing guesswork while designing (rather than discovering) antibodies. We replace the long, iterative process with faster, customizable, multi-parameter optimization.